01 Overview
Time-of-flight secondary-ion mass spectrometry (ToF-SIMS) gives a per-pixel mass spectrum — 129 ion channels in this case — at a 2 mm field of view from a stitched 1024×1024 SR2×2 HMR acquisition. The challenge is turning that hyperstack into something a dental clinician would recognize: a four-class anatomical map. Three approaches are compared and benchmarked against a clinician-traced reference.
- Channels: 129 ion peaks
- Resolution: 1024×1024 px, 2 mm FOV (stitched SR2×2 HMR)
- Anatomy classes: dentin, inner enamel, outer enamel, epoxy
02 Method
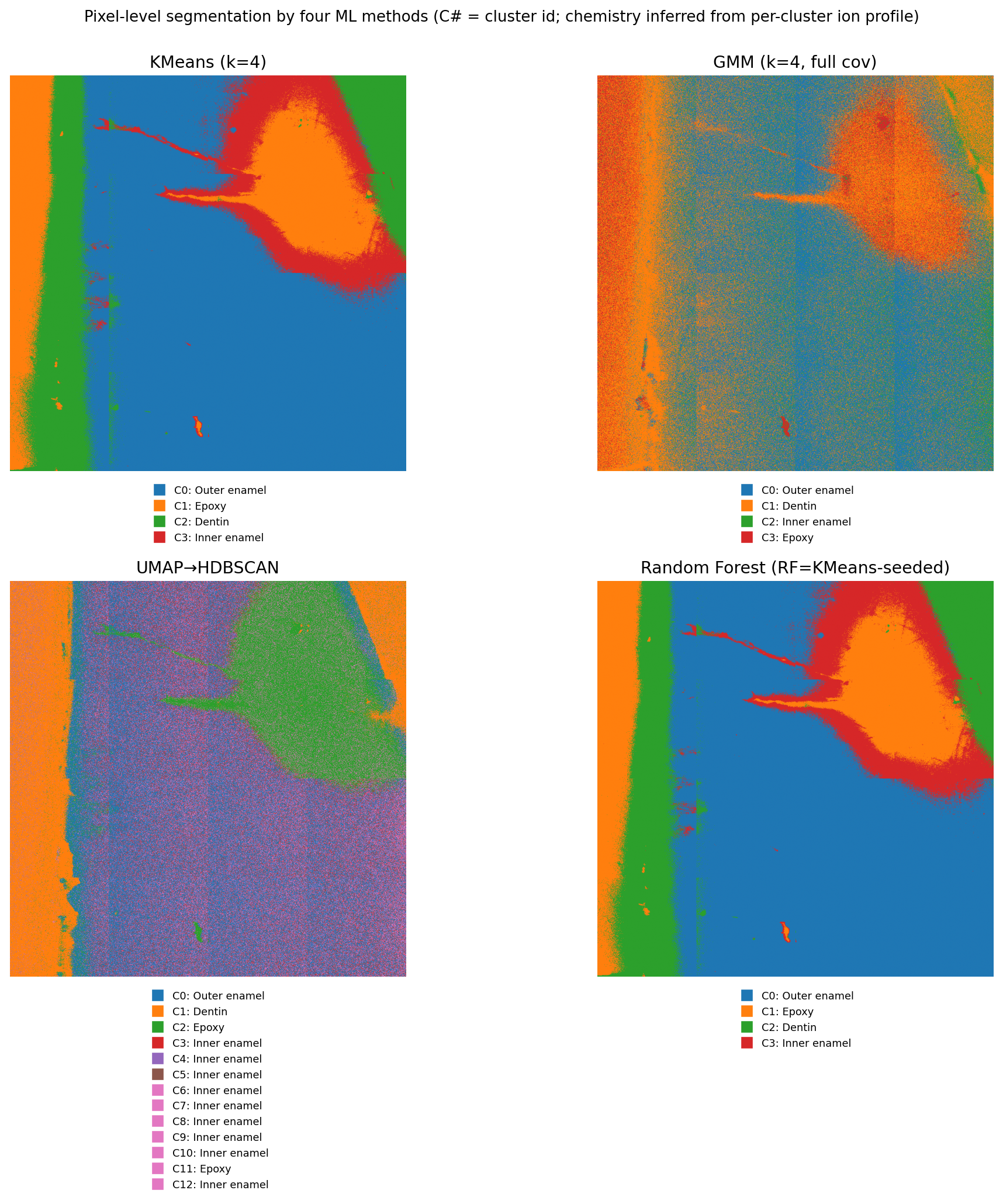
Three segmentation strategies were compared:
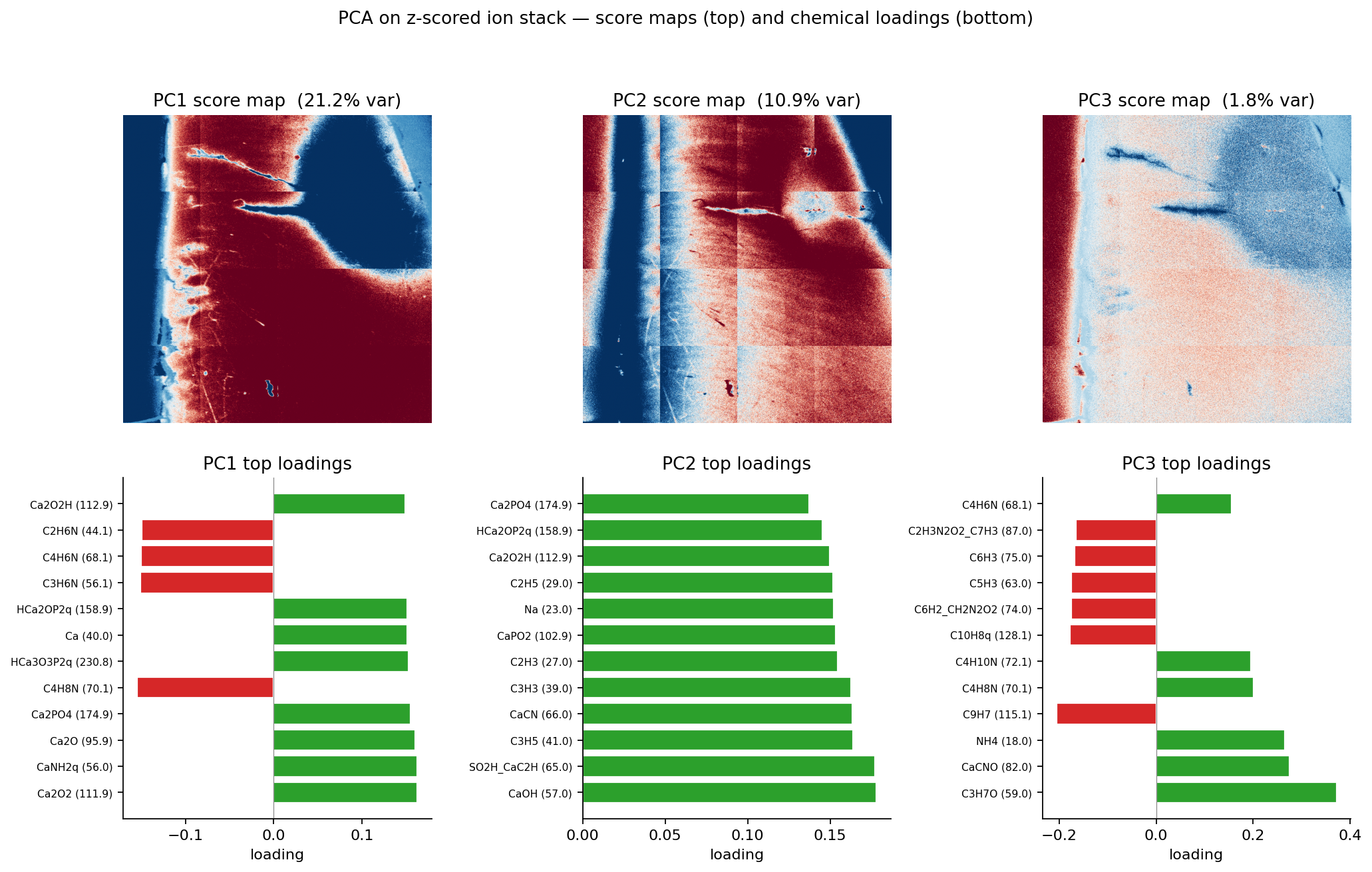
- Dan-style PCA + thresholds — three histogram-trough thresholds on the first three principal components. Best balance of accuracy and interpretability; chosen as the production method.
- Polygon-trained random forest / U-Net — supervised on clinician-drawn polygons; sensitive to acquisition variability
- HDBSCAN + NMF — fully unsupervised; useful for exploratory ion-class discovery
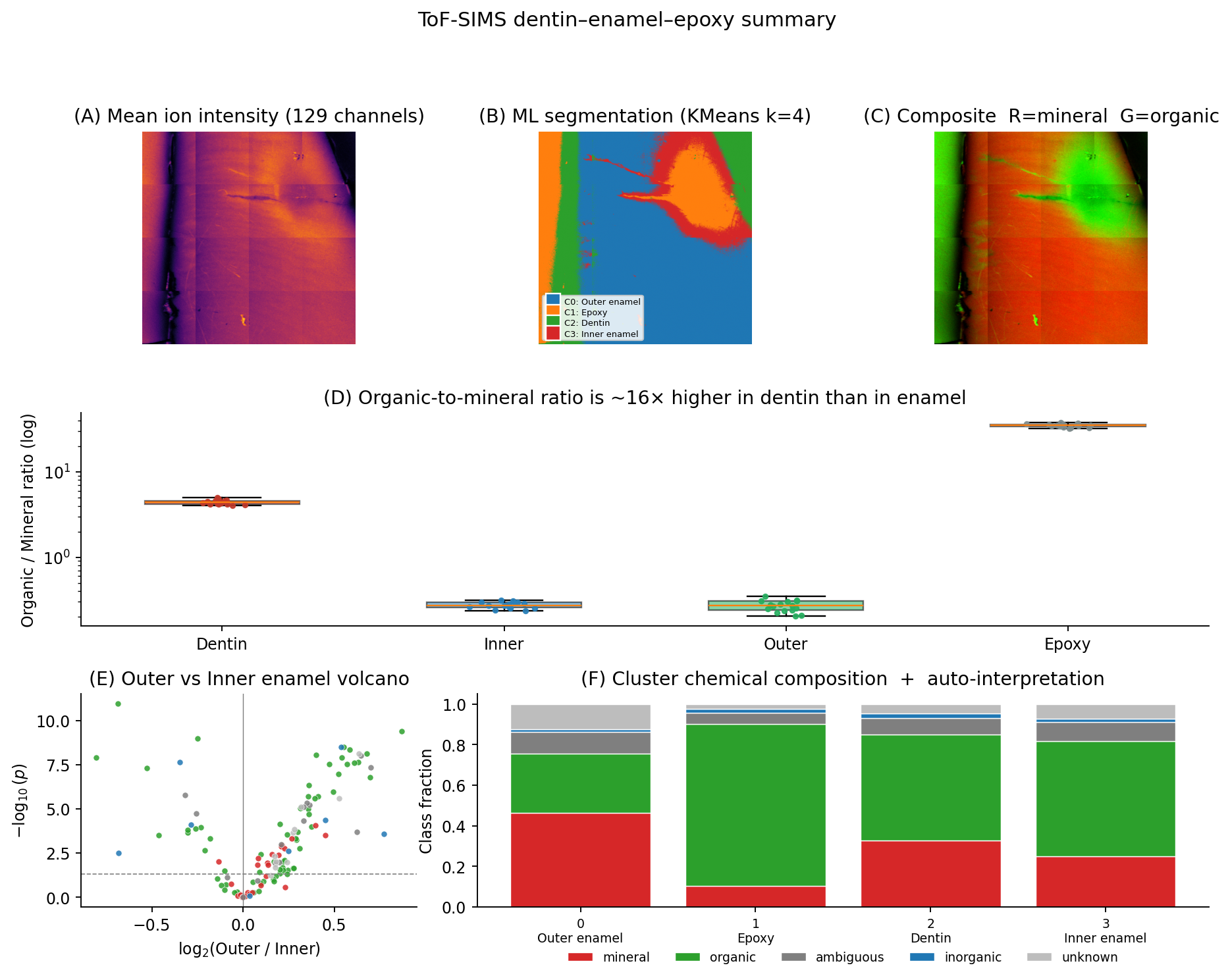
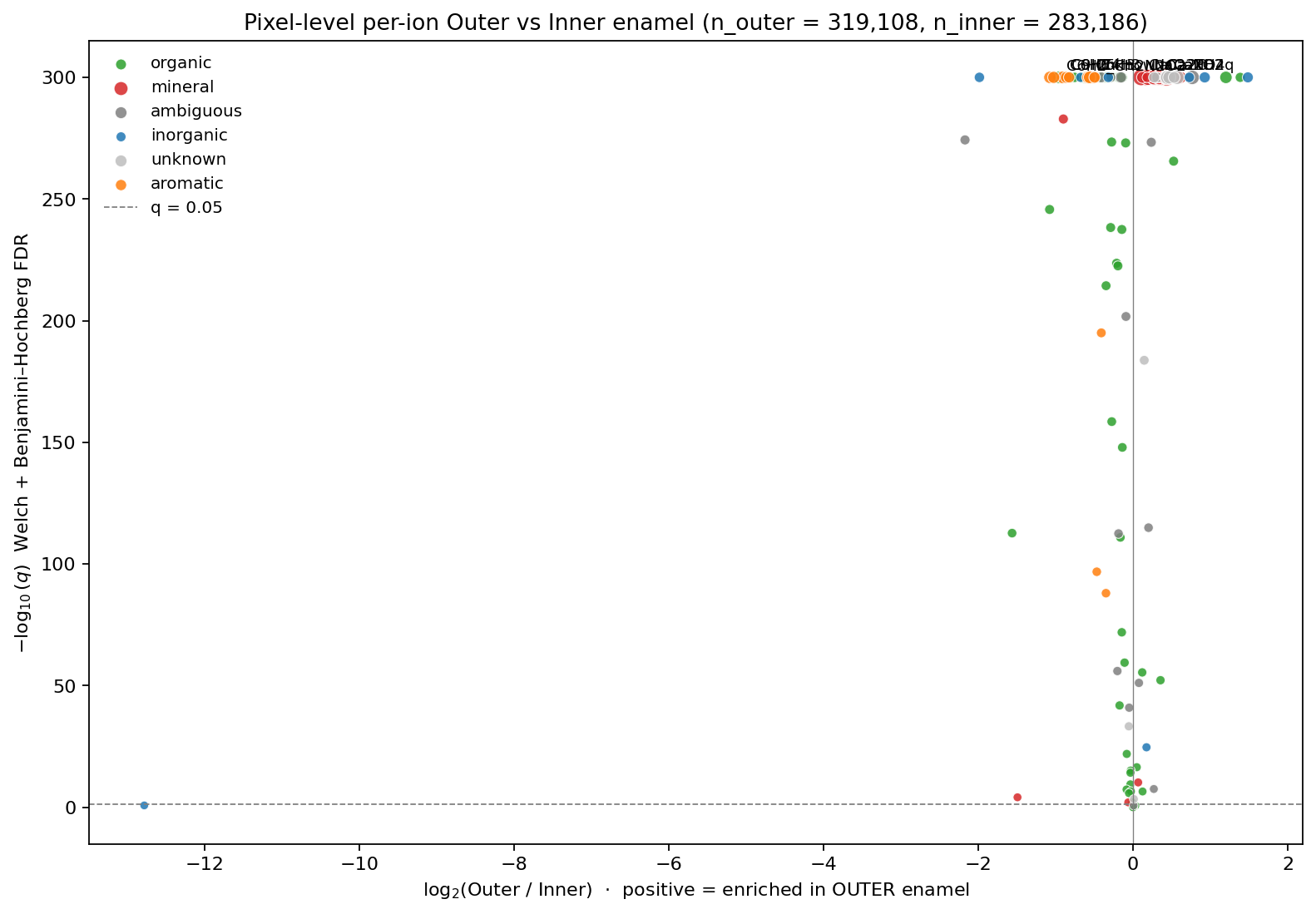
A per-ion outer-vs-inner compositional analysis (Welch t-test, Cohen's d, Benjamini-Hochberg multiple-correction) was then applied at the segmentation boundaries.
03 Visuals
04 Result
The mineral signal (Ca, P, hydroxyapatite fragments) agrees between outer and inner enamel — as expected from histology. The organic signal doesn't: outer enamel shows systematically different distributions for several CN-, CHN-, and protein-fragment ions, pointing to organic-matrix gradients across the enamel that aren't visible in conventional micro-anatomy. That shift is the publishable finding from this dataset.